| Abstract: | 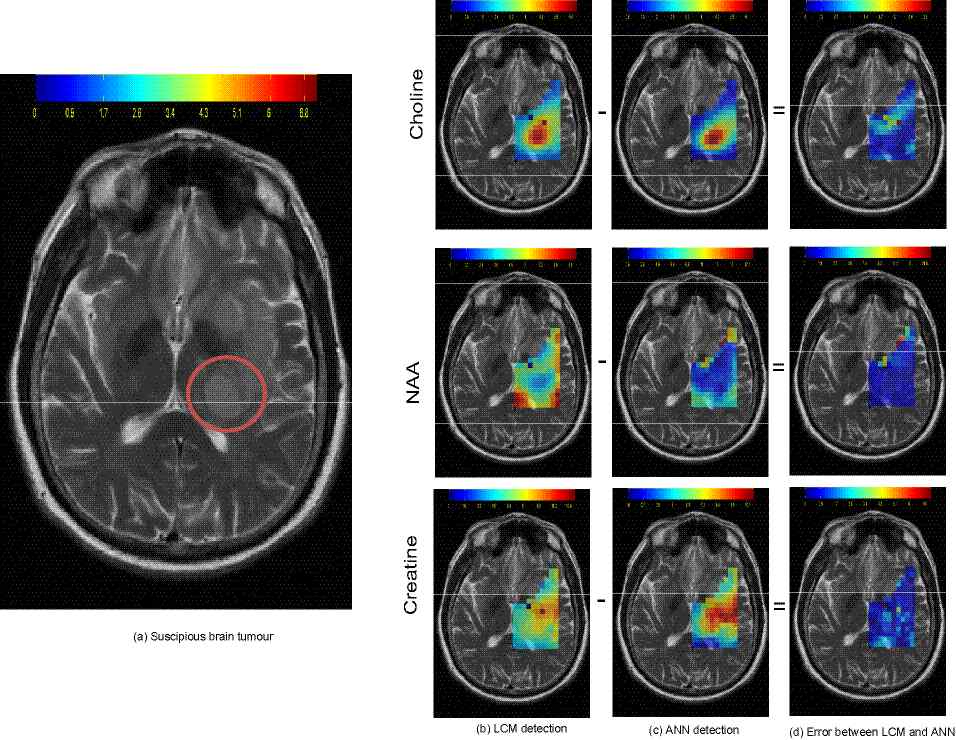 Magnetic resonance has proven to be a successful method of in-vivo imaging. Although
MRI can help detect various pathologies, its ability to classify the nature of the pathological
tissue is limited. Magnetic resonance spectroscopy allows identifying metabolite
content of the tissue and estimating the metabolite concentration. Map of metabolite concentration
along with the MR image allows proper classification of many pathologies, for
example progressive tumorous tissue identification in human brain. Standard methods
used to analyze nuclear magnetic resonance spectra such as singular value decomposition
or curve fitting algorithms are very time consuming taking several minutes to analyze
spectrum from a single voxel. To analyze the spectra from a chemical shift imagine sequence
(CSI) in maximal resolution hundreds of spectra need to be processed. The suggested
ANN framework proved to be much faster. Networks were trained on the outputs
of LCModel curve fitting algorithm. Time needed to process a spectrum from a single
voxel was reduced to the order of seconds. The total time needed to analyze a CSI in full
resolution (hundreds of spectra) was significantly reduced to 5 minutes. Magnetic resonance has proven to be a successful method of in-vivo imaging. Although
MRI can help detect various pathologies, its ability to classify the nature of the pathological
tissue is limited. Magnetic resonance spectroscopy allows identifying metabolite
content of the tissue and estimating the metabolite concentration. Map of metabolite concentration
along with the MR image allows proper classification of many pathologies, for
example progressive tumorous tissue identification in human brain. Standard methods
used to analyze nuclear magnetic resonance spectra such as singular value decomposition
or curve fitting algorithms are very time consuming taking several minutes to analyze
spectrum from a single voxel. To analyze the spectra from a chemical shift imagine sequence
(CSI) in maximal resolution hundreds of spectra need to be processed. The suggested
ANN framework proved to be much faster. Networks were trained on the outputs
of LCModel curve fitting algorithm. Time needed to process a spectrum from a single
voxel was reduced to the order of seconds. The total time needed to analyze a CSI in full
resolution (hundreds of spectra) was significantly reduced to 5 minutes.
|
|---|
 Magnetic resonance has proven to be a successful method of in-vivo imaging. Although
MRI can help detect various pathologies, its ability to classify the nature of the pathological
tissue is limited. Magnetic resonance spectroscopy allows identifying metabolite
content of the tissue and estimating the metabolite concentration. Map of metabolite concentration
along with the MR image allows proper classification of many pathologies, for
example progressive tumorous tissue identification in human brain. Standard methods
used to analyze nuclear magnetic resonance spectra such as singular value decomposition
or curve fitting algorithms are very time consuming taking several minutes to analyze
spectrum from a single voxel. To analyze the spectra from a chemical shift imagine sequence
(CSI) in maximal resolution hundreds of spectra need to be processed. The suggested
ANN framework proved to be much faster. Networks were trained on the outputs
of LCModel curve fitting algorithm. Time needed to process a spectrum from a single
voxel was reduced to the order of seconds. The total time needed to analyze a CSI in full
resolution (hundreds of spectra) was significantly reduced to 5 minutes.
Magnetic resonance has proven to be a successful method of in-vivo imaging. Although
MRI can help detect various pathologies, its ability to classify the nature of the pathological
tissue is limited. Magnetic resonance spectroscopy allows identifying metabolite
content of the tissue and estimating the metabolite concentration. Map of metabolite concentration
along with the MR image allows proper classification of many pathologies, for
example progressive tumorous tissue identification in human brain. Standard methods
used to analyze nuclear magnetic resonance spectra such as singular value decomposition
or curve fitting algorithms are very time consuming taking several minutes to analyze
spectrum from a single voxel. To analyze the spectra from a chemical shift imagine sequence
(CSI) in maximal resolution hundreds of spectra need to be processed. The suggested
ANN framework proved to be much faster. Networks were trained on the outputs
of LCModel curve fitting algorithm. Time needed to process a spectrum from a single
voxel was reduced to the order of seconds. The total time needed to analyze a CSI in full
resolution (hundreds of spectra) was significantly reduced to 5 minutes.